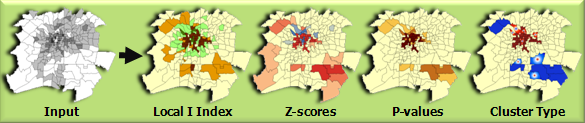
This tool creates a new Output Feature Class with the following attributes for each feature in the Input Feature Class: Local Moran's I index, z-score, pseudo p-value, and cluster/outlier type (COType).
The z-scores and p-values are measures of statistical significance which tell you whether or not to reject the null hypothesis, feature by feature. In effect, they indicate whether the apparent similarity (a spatial clustering of either high or low values) or dissimilarity (a spatial outlier) is more pronounced than one would expect in a random distribution. The p-values and z-scores in the Output Feature Class do not reflect any FDR (False Discovery Rate) corrections.
A high positive z-score for a feature indicates that the surrounding features have similar values (either high values or low values). The COType field in the Output Feature Class will be HH for a statistically significant cluster of high values and LL for a statistically significant cluster of low values.
A low negative z-score (for example, less than -3.96) for a feature indicates a statistically significant spatial data outlier. The COType field in the Output Feature Class will indicate if the feature has a high value and is surrounded by features with low values (HL) or if the feature has a low value and is surrounded by features with high values (LH).
The COType field will always indicate statistically significant clusters and outliers for a 95 percent confidence level. Only statistically significant features have values for the COType field. When you check the optional Apply False Discovery Rate (FDR) Correction parameter, statistical significance is based on a corrected 95 percent confidence level.
Default rendering for the Output Feature Class is based on the values in the COType field.
The output of this tool also includes a histogram charting the value of the Input Field and a Moran's scatterplot. These charts can be accessed from the Contents pane under the Output Feature Class.
Permutations are used to determine how likely it would be to find the actual spatial distribution of the values you are analyzing. For each permutation, the neighborhood values around each feature are randomly rearranged and the Local Moran's I value calculated. The result is a reference distribution of values that is then compared to the actual observed Moran's I to determine the probability that the observed value could be found in the random distribution. The default is 499 permutations; however, the random sample distribution is improved with increasing permutations, which improves the precision of the pseudo p-value.
If the Number of Permutations parameter is set to 0, the result is a traditional p-value instead of a pseudo p-value and the z-score is based on the randomization null hypothesis computation. For more information on z-scores and p-values, see What is a z-score? What is a p-value?
When the Input Feature Class is not projected (that is, when coordinates are given in degrees, minutes, and seconds) or when the output coordinate system is set to a Geographic Coordinate System, distances are computed using chordal measurements. Chordal distance measurements are used because they can be computed quickly and provide very good estimates of true geodesic distances, at least for points within about 30 degrees of each other. Chordal distances are based on an oblate spheroid. Given any two points on the earth's surface, the chordal distance between them is the length of a line, passing through the three-dimensional earth, to connect those two points. Chordal distances are reported in meters.
Caution:
Be sure to project your data if your study area extends beyond 30 degrees. Chordal distances are not a good estimate of geodesic distances beyond 30 degrees.
When chordal distances are used in the analysis, the Distance Band or Threshold Distance parameter, if specified, should be given in meters.
-
For line and polygon features, feature centroids are used in distance computations. For multipoints, polylines, or polygons with multiple parts, the centroid is computed using the weighted mean center of all feature parts. The weighting for point features is 1, for line features is length, and for polygon features is area.
The Input Field should contain a variety of values. The math for this statistic requires some variation in the variable being analyzed; it cannot solve if all input values are 1, for example. If you want to use this tool to analyze the spatial pattern of incident data, consider aggregating your incident data. The Optimized Hot Spot Analysis tool may also be used to analyze the spatial pattern of incident data.
Incident data are points representing events (crime, traffic accidents) or objects (trees, stores) where your focus is on presence or absence rather than some measured attribute associated with each point.
Your choice for the Conceptualization of Spatial Relationships parameter should reflect inherent relationships among the features you are analyzing. The more realistically you can model how features interact with each other in space, the more accurate your results will be. Recommendations are outlined in Selecting a Conceptualization of Spatial Relationships: Best Practices. Here are some additional tips:
- Fixed distance band
Uses a Distance Band or Threshold Distance and it will ensure each feature has at least one neighbor. This is important, but often this default will not be the most appropriate distance to use for your analysis. Additional strategies for selecting an appropriate scale (distance band) for your analysis are outlined in Selecting a fixed distance band value.
- Inverse distance or Inverse distance squared
When zero is entered for the Distance Band or Threshold Distance parameter, all features are considered neighbors of all other features; when this parameter is left blank, the default distance will be applied.
Weights for distances less than 1 become unstable when they are inverted. Consequently, the weighting for features separated by less than 1 unit of distance are given a weight of 1.
For the inverse distance options (Inverse distance, Inverse distance squared, or Zone of indifference), any two points that are coincident will be given a weight of 1 to avoid zero division. This assures features are not excluded from analysis.
Additional options for the Conceptualization of Spatial Relationships parameter, including space-time relationships, are available using the Generate Spatial Weights Matrix tool. To take advantage of these additional options, construct a spatial weights matrix file prior to analysis; select Get spatial weights from file for the Conceptualization of Spatial Relationships parameter; and for the Weights Matrix File parameter, specify the path to the spatial weights file you created.
-
More information about space-time cluster analysis is provided in the Space-Time Analysis documentation.
-
Map layers can be used to define the Input Feature Class. When using a layer with a selection, only the selected features are included in the analysis.
If you provide a Weights Matrix File with a .swm extension, this tool is expecting a spatial weights matrix file created using the Generate Spatial Weights Matrix tool; otherwise, this tool is expecting an ASCII-formatted spatial weights matrix file. In some cases, behavior is different depending on which type of spatial weights matrix file you use:
- ASCII-formatted spatial weights matrix files:
- Weights are used as is. Missing feature-to-feature relationships are treated as zeros.
- If the weights are row standardized, results will likely be incorrect for analyses on selection sets. If you need to run your analysis on a selection set, convert the ASCII spatial weights file to an SWM file by reading the ASCII data into a table, then using the Convert table option with the Generate Spatial Weights Matrix tool.
- SWM-formatted spatial weights matrix file:
- If the weights are row standardized, they will be restandardized for selection sets; otherwise, weights are used as is.
Running your analysis with an ASCII-formatted spatial weights matrix file is memory intensive. For analyses on more than 5,000 features, consider converting your ASCII-formatted spatial weights matrix file into an SWM-formatted file. First, put your ASCII weights into a formatted table (using Excel, for example). Next, run the Generate Spatial Weights Matrix tool using Convert table for the Conceptualization of Spatial Relationships parameter. The output will be an SWM-formatted spatial weights matrix file.
The Output Feature Class is automatically added to the table of contents with default rendering applied to the COType field. The rendering applied is defined by a layer file in <ArcGIS Pro>\Resources\ArcToolBox\Templates\Layers. You can reapply the default rendering, if needed, by using the Apply Symbology From Layer tool.
-
The Output Feature Class includes a SOURCE_ID field which allows you to Join it to the Input Feature Class, if needed.
-
The Modeling Spatial Relationships help topic provides additional information about this tool's parameters.
Caution:
When using shapefiles, keep in mind that they cannot store null values. Tools or other procedures that create shapefiles from nonshapefile inputs may store or interpret null values as zero. In some cases, nulls are stored as very large negative values in shapefiles. This can lead to unexpected results. See Geoprocessing considerations for shapefile output for more information.
When using this tool in Python scripts, the result object returned from tool execution has the following outputs:
| Position | Description | Data Type |
|---|
0 | Output feature class | Feature Class |
1 | Index field name | Field |
2 | ZScore field name | Field |
3 | Probability field name | Field |
4 | COType field name | Field |
5 | Source ID field name | Field |